Why do we use 2 restriction enzymes
The use of 2 different enzymes makes self ligation of the vector impossible and makes the insertion unidirectional. Whereas in the case of single digest, selfligation occurs and insertion may occur in both ways. Overall the use of 2 RE increases the probability to get the right construct.
Why are two restriction enzymes used in gel electrophoresis?
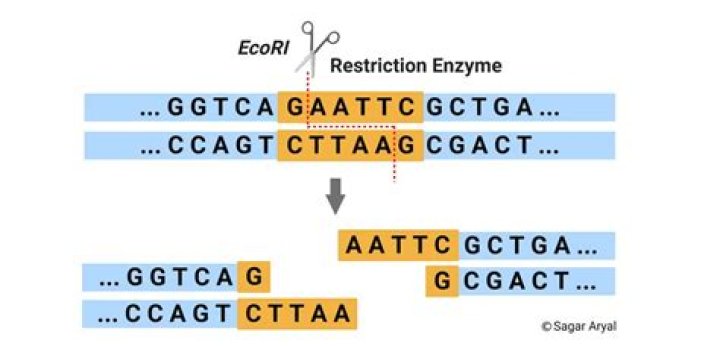
These enzymes cut both strand of the target DNA at different spots creating 3‘- or 5’-overhangs of 1 to 4 nucleotides (so-called sticky ends). To be able to clone a DNA insert into a cloning or expression vector, both have to be treated with two restriction enzymes that create compatible ends.
Why are Type 2 restriction enzymes used in cloning?
Type II restriction enzymes are the familiar ones used for everyday molecular biology applications such as gene cloning and DNA fragmentation and analysis. These enzymes cleave DNA at fixed positions with respect to their recognition sequence, creating reproducible fragments and distinct gel electrophoresis patterns.
Why are the same two restriction enzymes used for both plasmids?
Restriction enzymes cut at specific sequences so the same restriction enzyme must be used because it will produce fragments with the same complementary sticky ends, making it possible for bonds to form between them.What is an advantage of using two restriction enzymes to cut a plasmid?
Then, you guessed it, use restriction enzymes. Restriction enzyme cloning takes advantage of the site specificity of these enzymes. The enzymes only cut (or “digest”) at specific DNA sequences —usually plasmid DNA in cloning. This specificity allows you to insert or ligate another piece of DNA at those sites.
Why is it important that the same enzyme or enzymes be used to cut both the plasmid and the insulin gene from the human DNA?
Why is it important that the same enzyme or enzymes be used to cut both the plasmid and the insulin gene from the human DNA? it is important to use the same enzyme so that both the ends of the insulin and plasmid connect. Which antibiotic would you use to determine if the recombinant DNA was taken in? Kanamycin.
What are the two types of restriction enzymes?
Today, scientists recognize three categories of restriction enzymes: type I, which recognize specific DNA sequences but make their cut at seemingly random sites that can be as far as 1,000 base pairs away from the recognition site; type II, which recognize and cut directly within the recognition site; and type III, …
Which enzyme is responsible for ligation of the two strands together?
DNA ligase is the protein responsible for linking, or ligating, Okazaki fragments together in order to form a single complete DNA strand.Why the same restriction enzyme must be used to extract the gene and open the loop of DNA in the bacterium?
Restriction endonuclease identifies and cuts the same pallindromic sequence in both DNA and Vector due to which when they will be mixed , their complementary bases will join and it will form the r-DNA, If both are cut with different RE , then on mixing they wont ligate with each other as their bases will not match.
Why are Type 2 restriction endonucleases used in recombinant DNA technology?Type II restriction enzymes have two properties useful in recombinant DNA technology. First, they cut DNA into fragments of a size suitable for cloning. Second, many restriction enzymes make staggered cuts generating single-stranded ends conducive to the formation of recombinant DNA.
Article first time published onWhy are Type 2 restriction endonucleases most useful in recombinant DNA technology?
Type II restriction endonucleases always cleave at or near their recognition sites. They produce small, well-defined fragments of DNA that help to characterize genes and genomes and that produce recombinant DNAs. Fragments of DNA produced by restriction endonucleases can be moved from one organism to…
Why are the restriction sites of Type II enzymes usually palindromic?
Explanation: Enzymes such as restriction enzymes have to recognize a very specific sequence in order to carry out its task. It binds to the DNA only in one specific configuration. … A palindromic sequence also increases the chance that both strands of DNA are cut.
How many restriction enzymes should be used?
In general, we recommend 5–10 units of enzyme per µg DNA, and 10–20 units for genomic DNA in a 1 hour digest.
Why do we most commonly used type 2 restriction endonucleases in cloning as opposed to the other types of restriction endonucleases?
Type II restriction enzymes are most commonly used for molecular biology applications, as they recognize stereotypical sequences and produce a predictable cleavage pattern.
Where do Type 2 restriction enzymes cut?
Type IIS enzymes generally bind to DNA as monomers and recognize asymmetric DNA sequences. They cleave outside of this sequence, within one to two turns of the DNA. By convention, the recognition sequence is written in the orientation in which cleavage occurs downstream, to the right of the sequence.
What is the purpose of restriction fragment?
Restriction Fragment Length Polymorphism (RFLP) These are bacterial enzymes used by scientists to cut DNA molecules at known locations. RFLPs (pronounced “rif lips”) are used as markers on genetic maps. Typically, gel electrophoresis is used to visualize RFLPs.
What is the advantage of using multiple restriction enzymes to cut the DNA during DNA fingerprinting?
enzymes to cut the DNA during DNA fingerprinting? Multiple sets of data provide more evidence than a single set and allow us to make stronger conclusions.
Why was it important to find an enzyme that would cut the plasmid at only one site?
1. Why was it important to find an enzyme that would cut the plasmid at only one site? … If the enzymes cut at multiple spots, then you would get multiple fragments.
Do you think restriction enzymes would be used to cut DNA from other organisms?
Restriction enzymes dismantle foreign DNA by cutting it into fragments. This disassembling process is called restriction. Recombinant DNA technology relies on restriction enzymes to produce new combinations of genes.
Why Restriction enzymes are used in genetic engineering?
The type II restriction enzymes are very specific to recognize and cut the desired DNA segments. … So, from the above discussion, we can conclude that Restriction enzymes are used in genetic engineering because they cut at specific recognition sequences of base pairs in DNA. Thus, option B) is the right answer.
Why Restriction enzymes are used in the process of genetic engineering?
The main steps of genetic engineering: Restriction enzymes are used to isolate the required gene from the chromosome . They cut the DNA at a specific sequence. Restriction enzymes leave sticky ends that are overhangs of DNA.
What role do Restriction enzymes play in genetic engineering?
A restriction enzyme is an enzyme isolated from bacteria that cuts DNA molecules at specific sequences. The isolation of these enzymes was critical to the development of recombinant DNA (rDNA) technology and genetic engineering.
Why do we need DNA ligase?
DNA ligases are critical DNA replication and repair enzymes; they have been widely used in molecular biology and biotechnology applications, such as cloning and next-generation DNA sequencing [1, 2]. DNA ligases catalyze the joining of adjacent 3′-hydroxyl and 5′-phosphorylated DNA termini in duplex DNA.
What enzyme joins two substrates?
In biochemistry, a ligase is an enzyme that can catalyze the joining (ligation) of two large molecules by forming a new chemical bond.
What 2 things does DNA polymerase do during DNA replication?
The DNA polymerases are enzymes that create DNA molecules by assembling nucleotides, the building blocks of DNA. These enzymes are essential to DNA replication and usually work in pairs to create two identical DNA strands from one original DNA molecule.
How is DNA digested by restriction endonuclease enzymes?
Restriction digestion is accomplished by incubation of the target DNA molecule with restriction enzymes – enzymes that recognize and bind specific DNA sequences and cleave at specific nucleotides either within the recognition sequence or outside of the recognition sequence.
Which ion is required for activity of type 2 restriction endonuclease?
The ion required for the activity of the type II restriction enzyme is Mg2+. Restriction enzymes or restriction endonucleases bind to a specific double-stranded DNA sequence and cleave the DNA at a specific site.
What is the importance of palindromic sequence?
Certain DNA sequences have a twofold inverted symmetry and are called self-complementary or palindromic sequences. Palindromic sequences are the usual recognition sites for restriction enzymes and frequently occur as essential elements in regulatory regions.
Why is restriction site important?
A restriction site is a sequence of approximately 6–8 base pairs of DNA that binds to a given restriction enzyme. These restriction enzymes, of which there are many, have been isolated from bacteria. Their natural function is to inactivate invading viruses by cleaving the viral DNA.
What is the benefit of utilizing multiple restriction enzymes for scientific applications?
if a researcher cuts dna from 2 different sources using the same restriction enzyme, it will allow them to insert a section of dna from one source to another. the recombined DNA plasmid is inserted into the bacteria.
What are Type 1 restriction enzymes used for?
Type I enzymes are complex, multisubunit, combination restriction-and-modification enzymes that cut DNA at random far from their recognition sequences. Originally thought to be rare, we now know from the analysis of sequenced genomes that they are common.